EZ-Epitope™
EZ-Epitope™ Profiler Free to download and use.
New Version 3.0 (March 16, 2026)
EZ-Epitope™ Performs prediction, ranking, and profiling of linear B-cell amino acid sequences.
Microsoft Windows 64-bit Console edition
In the immune system, B cells play a central role in initiating humoral immune responses. They achieve this by specifically recognizing particular regions on antigen molecules through their surface B cell receptors (BCRs). This recognition triggers B cell activation, proliferation, and differentiation into plasma cells, which ultimately produce and secrete large quantities of highly specific antibodies. These antibodies subsequently bind to specific sites on the antigen, known as antibody epitopes, to neutralize pathogens or toxins. Epitope mapping is the process of experimentally identifying the binding site, or epitope, of an antibody on its target antigen (usually, on a protein). Identification and characterization of antibody binding sites aid in the discovery and development of new therapeutics, vaccines, and diagnostics.
B Cell Epitope Properties:
An antigen is any substance that the immune system can recognize as being foreign and which provokes an immune response. Since antigens are usually proteins that are too large to bind as a whole to any receptor, only specific segments that form the antigen bind with a specific antibody. Such segments are called epitopes. A B cell epitope is a specific region on an antigen molecule that is recognized and bound by a BCR. This interaction initiates the humoral immune response by activating B cells. B-cell epitopes consists of a hydrophobic core flanked by charged amino acids.
Key properties:
- Sequence-Based Methods - These methods rely solely on the amino acid sequence of the antigen. They often use statistical correlations with known linear epitopes, looking for regions enriched in properties like hydrophilicity, flexibility or coil.
- Linear vs. Conformational - While conformational epitopes are the typical form of B cell epitopes recognized on native antigens by BCRs, in some cases, B cells can also recognize linear epitopes, which consist of a continuous sequence of amino acids. This often occurs after the antigen has been processed by antigen-presenting cells (APCs), and linear peptide fragments are presented and recognized by B cells.
- A linear epitope (also sequential epitope) is an epitope - a binding site on an antigen that is recognized by antibodies by its linear sequence of amino acids (i.e. primary structure).
- Accessibility and Surface Exposure - B cell epitopes must be located on the surface of the antigen, making them physically accessible for recognition and binding by the cell-surface BCR.
- Flexibility - Appropriate flexibility can potentially facilitate the B cell epitope in adapting to the binding conformation of the BCR or antibodies.
- Charge and Hydrophilicity - B cell epitopes often reside in hydrophilic regions on the protein surface, facilitating interaction with the aqueous environment and the binding sites of BCRs and antibodies. Charged residues can play a significant role in specific electrostatic interactions.
EZ-Epitope profiler:
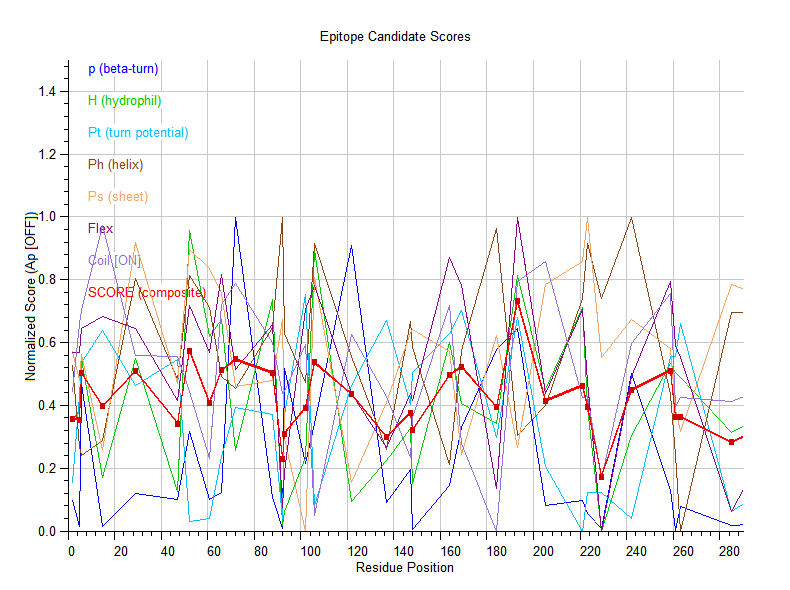
Generates amino acid property profiles along a protein sequence using a sliding window average. It provides 7 different physicochemical scales spanning hydrophobicity, coil structure propensity, antigenic propensity, flexibility,, beta-turn potential, helix potential, sheet potential, and beta-turn probability. The scale profiler unifies all major amino acid property scales into one comprehensive final scale allowing the comparison of different properties for the same amino acid sequence. The scoring combines multiple biophysical signals into antigenic scores:
- P (beta-turn probability): higher = likely to be in a loop/exposed region
- H (hydrophilicity): higher = more surface-exposed
- Pt (beta-turn potential): values >1.0 favor turn conformation
- Ps, Ph: sheet/helix potentials — lower = better (buried in secondary structure)
- C (conformation): t/t* = best, - = ok, s/h = less surface exposure
- Average flexibility (Bhaskaran & Ponnuswamy)
- Propensity for random-coil conformation (Deleage & Roux)
- Surface exposure frequencies (Janin & Wodak)
- Antigenic propensity for surface exposure (Kolaskar & Tongaonkar)
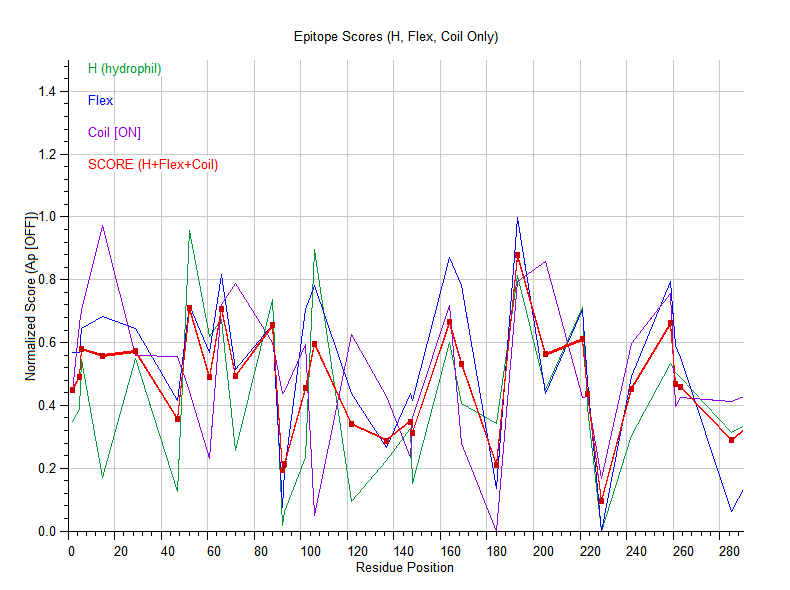
- Balances coil-rich loops with hydrophilic regions
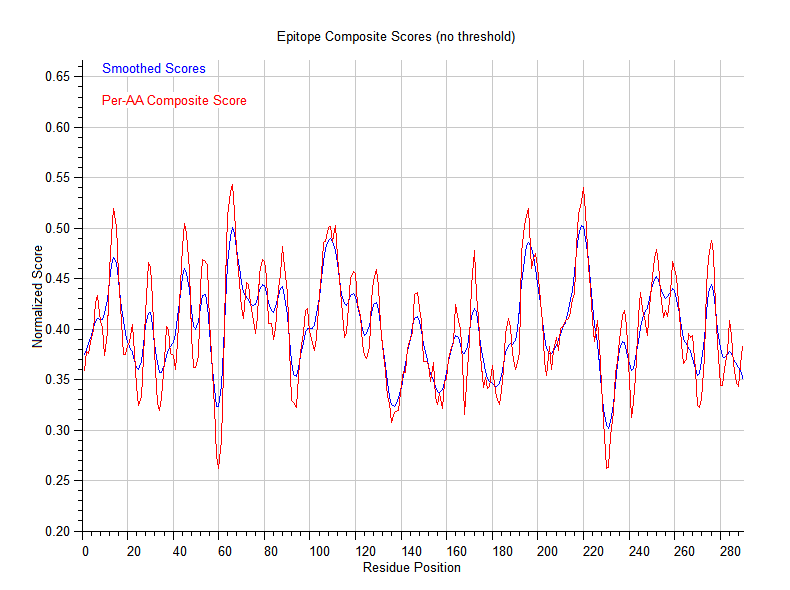
- Computes a per-residue antigenic score profile over the entire sequence
- Prediction, ranking, and per-residue antigenic profile
Reference:
- Chou PY, Fasman GD (1974). Conformational parameters for amino acids in helical, beta-sheet, and random coil regions calculated from proteins. Biochemistry 13:211–222.
- Janin J, Wodak S (1978). Conformation of Amino Acid Side-Chains in Proteins. J. Mol. Biol. 125:357-386.
- Hopp TP, Woods KR (1981). Prediction of protein antigenic determinants from amino acid sequences. Proc. Natl. Acad. Sci. USA 78:3824–3828.
- Hopp TP, Woods KR (1983). A computer program for predicting protein antigenic determinants. Molecular Immunology 20:483-489.
- Hopp TP (1986). Protein surface analysis: Methods for identifying antigenic determinants. J. Immunol. Methods 88:1–18.
- Hopp TP, Woods KR (1987). A computer program for predicting protein antigenic determinants. Anal. Biochem. 165:200–207.
- Welling GW et al. (1985). Prediction of sequential antigenic regions in proteins. FEBS Lett. 188:215–218.
- Krchnák V, Mach O, Malý A (1987). Computer prediction of potential immunogenic determinants from protein amino acid sequence. Anal. Biochem. 165:200–207.
- Deléage G and Roux B (1987). An algorithm for protein secondary structure prediction based on class prediction. Protein Engineering 1 (4):289–294.
- Jameson BA, Wolf H (1988). The antigenic index: a novel algorithm for predicting antigenic determinants. Comput. Appl. Biosci. 4:181–186.
- Bhaskaran R, Ponnuswamy PK (1988). Positional flexibilities of amino acid residues in globular proteins. Int. J. Peptide Protein Res. 32, 241-255.
- Kolaskar, AS and Tongaonkar, PC (1990). A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett. 276, 172-174.
- Kringeluma JV, Nielsena M, Padkjærb SB, and Lund O (2013). Structural analysis of B-cell epitopes in antibody:protein. complexes. Mol Immunol. 53(1-2):24–34.
- Andersen PH, Nielsen M, Lund O (2006). Prediction of residues in discontinuous B-cell epitopes using protein 3D structures. Protein Sci. 15:2558–2567.
- Ofran Y, Schlessinger A, Rost B (2008). Automated identification of complementarity determining regions (CDRs) reveals peculiar characteristics of CDRs and B cell epitopes. J. Immunol. 181:6230-6235.
- Rubinstein ND, Mayrose I, Halperin D, Yekutieli D, Gershoni JM, Pupko T (2008). Computational characterization of B-cell epitopes. Mol. Immunol. 45:3477–3489.
- Zhao L, Li J (2010) Mining for the antibody-antigen interacting associations that predict the B cell epitopes. BMC Struct. Biol. 10(Suppl 1):S6.
- Sun J, Xu T, Wang S, Li G, Wu D, Cao Z (2011). Does difference exist between epitope and non-epitope residues? Analysis of the physicochemical and structural properties on conformational epitopes from B-cell protein antigens. Immunome Res. 7:1–11.
Download the Free to use software at the bottom of the page.
Consider donating to support the further development of software. Thank you.

Donate
Donate to support further development of software.
$
10
DONATE TO EZ-EPITOPE
EZ-Epitope predicts B-cell antigen ranking, profiling, amino acid conversion.
Finally, select 'Calculate to display the results'.
Download the latest version of EZ-Epitope™
EZ-Epitope predicts antigen ranking, profiling, amino acid conversion, and nucleotide complementary sequence.
o Developed by F.W. Perrella, Ph.D. (February 13, 2026)
o www.EnzymKinetics.com
"THERE IS NO WARRANTY, EXPRESSED OR IMPLIED, FOR THIS PROGRAM. ITS SUITABILITY FOR COMMERCIAL USE OR FOR ANY PARTICULAR PURPOSE IS NOT GUARANTEED."
Download the software zip file to a single folder, Unzip it and then run EZ-Epitope.exe
Download and unZip the file in a folder (e.g., C:\EZ-Epitope.zip\) on a computer you choose to license the software, and then run EZ-Epitope.exe. The lack of formal Windows installation is intentional, as it allows installation without administrative privileges (no dynamic link libraries (DLL) are used) and no network connection. EZ-Epitope runs in a MS Windows 64-bit Console.
Note: EZ-Epitope.zip has been scanned by Windows Defender.
File size: 587.58 KB
Download the EZ-Radioisotope-Decay ZIP file to your computer and unzip the file in a folder on your computer.
Note: If your browser flags the downloaded zip file with a cautionary warning, you have the option of selecting "keep" to continue the download process. The EZ-Epitope.zip file was scanned with Microsoft Windows Defender and shown not to have any detectable threats. The EZ-Epitope software does not send any information back to us or anyone else, and it does not require a network connection to run. It is a standalone executable file (no DLL files) .
Disclaimer of 'False Positive' virus: EZ-Epitope.exe is an independently distributed program, and as such it may be flagged as a 'false positive' by your antivirus/antimalware software. Therefore, it is advisable to create an exclusion for the EZ-Epitope.exe software to avoid the appearance of 'false positive' errors during execution.
Your cart is empty
0
item(s)
/
$0
Checkout
Clear cart