Nonlinear Regression Software for Curve-fitting Data to Kinetic Models
- BestCurvFit, Manchester, New Hampshire, USA, 03103
- drfrank88@gmail.com
EZ-SCAFIT™ (FREE)
EZ-SCAFIT™ Ligand Receptor Binding - Free to download and use.
New Version 2.0 (March 16, 2026)
EZ-SCAFIT™ Performs non-linear least squares curve-fitting of experimental receptor binding data.
Microsoft Windows 64-bit Console edition
EZ-SCAFIT performs the non-linear least square curve fit of experimental receptor binding data using estimates of KA, Rmax and Nonspecific. The Bound ligand vs Total ligand coordinate system is used in the analysis of one or two binding sites for saturation or competition assays.
EZ-SCAFIT was developed based on the SCAFIT methods originally developed by Peter J. Munson and David Rodbard at the U.S. National Institutes of Health (NIH) for analysis of ligand-receptor binding. LIGAND/SCAFIT software was authored and developed by Federal employees as a part of their official duties, it therefore constitutes a work of the U.S. Government under 17 U.S.C. section 105 and is in the public domain. The foundational work describing these methods appeared in the classic papers:
- 1. P.J. Munson and D. Rodbard, LIGAND: a versatile computerized approach for characterization of ligand-binding systems, Analytical Biochemistry 107, 220–239 (1980).
- 2. H.A. Feldman, Mathematical Theory of Ligand-Binding at Equilibrium Analytical Biochemistry 48, 317-338 (1972).
EZ-SCAFIT:
- Developed independently based on methodology similar to SCAFIT
- Uses the general binding model described by Feldman (1972) and implemented by Munson & Rodbard (1980)
- Adapted for 64-bit Microsoft Windows/Console
- Uses Nelder-Mead Simplex, Conjugate Gradient, and Levenberg-Marquardt optimization routines
- Uses weighted nonlinear least squares with numerical derivatives
- Implements Saturation and Competition displacement binding
- Reorders data files, Saturation (1st), Competition (next)
- Fits data to models containing one or two binding sites
- All data must be in Molar [M] units
- Varying hot ligand [M] must not be less than 1.0E-14 M
- Applies automatic initial parameter estimates (K, R, N)
- Allows refitting data using last fitted parameter values
- Implements bootstrap parameter standard errors
- Performs Runs test of curve-fit residual patterns
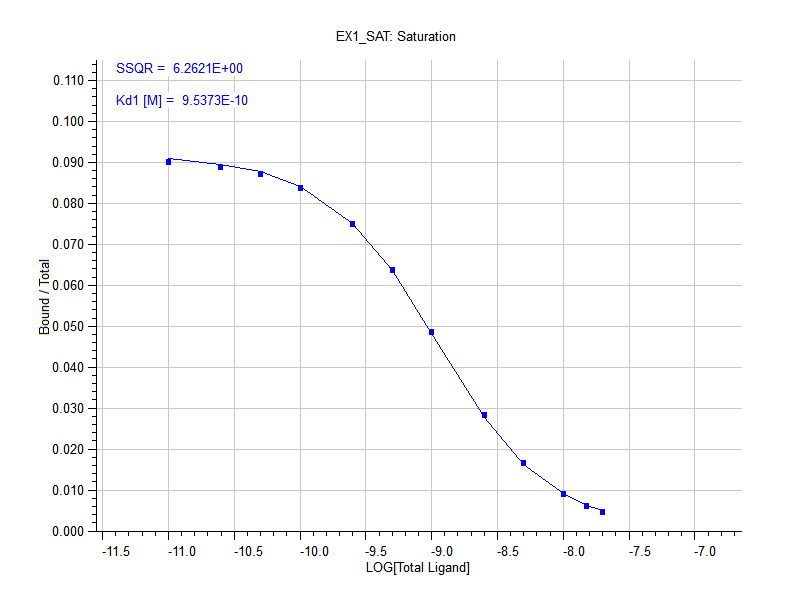
- Creates plots of Log(Total Ligand) versus Bound/Total Ligand
The EZ-SCAFIT program implements Feldmans 1972 framework by:
- Accepting experimental data (total ligand added, measured binding)
- Specifyng the model (number of sites, ligands, interactions)
- Solving the coupled equations iteratively for each data point
- Using weighted nonlinear least squares with numerical derivatives
- Outputs fitted parameters with standard errors and confidence intervals
- H.A. Feldman, Mathematical Theory of Ligand-Binding at Equilibrium. Analytical Biochemistry 48, 317-338 (1972).
This approach transforms ligand binding from a qualitative, graphical Scatchard analysis into a quantitative method with statistically justified parameter estimates.
Example: Two Ligands, One Receptor
Consider a competitive binding experiment with radioligand [L*] and unlabeled competitor [L] binding to receptor [R]:
Mass action:
[LR] = [L]free * [R]free / Kd1 [LR] = [L]free * [R]free / Kd2
Conservation:
[L*]total = [L*]free + [L*R] [L]total = [L]free + [LR] [R]total = [R]free + [L*R] + [LR]
Combining the mass action and conservation equations creates a system of coupled nonlinear algebraic equations that can be solved interatively to find all [Li]free and [Rj]free values that satisfy both mass action and conservation simultaneously.
The coupled system (3 equations, 3 unknowns):
[L*]total = [L*]free + ([L*]free * [R]free / Kd1)
[L]total = [L]free + ([L]free * [R]free / Kd2)
[R]total = [R]free + ([L*]free * [R]free / Kd1) + ([L]free * [R]free / Kd2)
Solving this system for [L*]free, [L]free, and [R]free allows the prediction of the measured radioligand binding [L*R] as a function of the competitor concentration [L]total. The curve-fitting of competition curves, allows the determination of both Kd1 and Kd2.
Feldmans approach can be equally explained using the association constant KA instead of the dissociation constant Kd. These constants are reciprocals of each other: KA = 1/Kd
- Kd (dissociation constant) has units of concentration (M) and represents the ligand concentration at which half the binding sites are occupied. A smaller Kd means higher affinity.
- KA (association constant) has units of inverse concentration (1/M) and represents the strength of the binding interaction. A larger KA means higher affinity.
Using KA, the conservation equations become:
For ligand i: [Li]total = [Li]free + SUMj{ (KA,ij * [Li]free * [Rj]free) }
For receptor type j: [Rj]total = [Rj]free + SUMi{ (KA,ij * [Li]free * [Rj]free) }
When comparing affinities, Kd = 1 nM binds tighter than 100 nM. When comparing reaction rates, KA = kon / koff
Receptor binding experiments are typically designed to measure the bound ligand using known total ligand concentrations. EZ-SCAFIT uses the general binding model described by Feldman (1972) and implemented by Munson & Rodbard (1980) to predict the bound and unbound ligand concentration.
- Feldman HA. Mathematical theory of complex ligand-binding systems at equilibrium. Analyt. Biochem. (1972); 48:317-338.
- Munson PI, Rodbard D. LIGAND: A versatile computerised approach for characterisation of ligand binding systems. Analyt. Biochem. (1980); 107:220-239.
There are two basic designs for receptor binding experiments:
- Saturation assays use increasing concentrations of the primary ligand (usually radioactive i.e. "hot") where the low concentrations are in the range of the Kd and high concentrations approach saturation of the binding site. The range of bound ligand concentration is typically measured over several orders of magnitude
- Displacement assays use a fixed concentration of a primary ligand (usually radioactive i.e. "hot") and increasing concentrations of a displacing ligand (usually non-radioactive i.e. "cold") to displace the primary ligand. The primary and displacing ligands can be isotopes of the same molecule or different molecules. The range of bound concentration is typically measuremented over one or two orders of magnitude.
"THERE IS NO WARRANTY, EXPRESSED OR IMPLIED, FOR THIS PROGRAM.
ITS SUITABILITY FOR COMMERCIAL USE OR FOR ANY PARTICULAR PURPOSE
IS NOT GUARANTEED."
Download the Free to use software at the bottom of the page.
Consider donating to support the further development of software. Thank you.

Donate
Donate to support further development of software.
$
10
DONATE
Your cart is empty
0
item(s)
/
$0
Checkout
Clear cart